
Matt McGuffie
Bioinformatics 2019

Bioinformatics 2019

Plasmids are extra-chromosomal, autonomously replicating segments of DNA found within bacteria and other organisms. In natural environments, they often provide bacteria with beneficial genes that allow for antibiotic resistance, heavy metal resistance, new metabolisms, pathogenesis, and more. These small pieces of DNA are relatively easy to modify in the lab and offer an easy solution for researchers attempting manipulate an organism’s genome. This makes the engineered plasmid is an invaluable tool in many fields of biology.
One of the first engineered plasmids, pBR322, has donated at least part of its sequence to over 92% of engineered plasmids in Addgene’s repository, making it the ancestor to the majority of engineered plasmids. However, even this “basic” plasmid has a complicated assembly history (see figure below1), comprised of a dozen or so distinct plasmids and transposons.
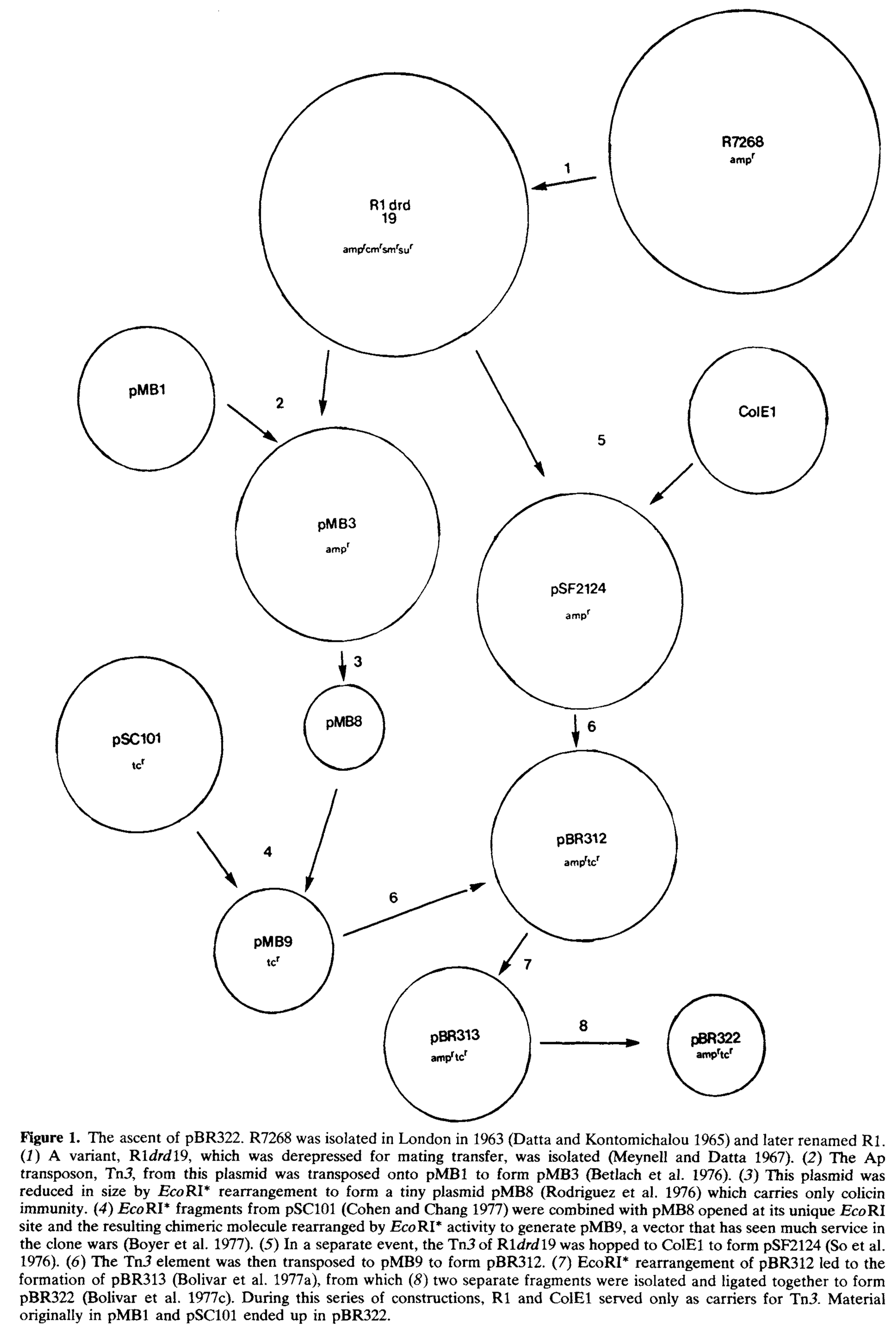
The various ancestral elements of pBR322 were assembled via cutting with restriction enzymes to obtain fragments which contained the desired function (antibiotic resistance, origin of replication, etc.), often taking DNA that has no explicit function along for the ride as well. This extra, not-explicitly-wanted DNA would then be carried into future plasmids constructed from pBR322 as well. Even with the advent of more precise cloning techniques, the construction of most plasmids follows a similar template of that of pBR322.
The inherited history of cloning scars litters engineered plasmids. The problems this extra DNA can create range from taking up valuable real estate on the plasmid, to making unintended and potentially toxic peptide fragments.
While current DNA computer aided design tools such as Geneious or Snapgene offer “auto-annotation” features, these programs only highlight features that are found in their entirety. Here I offer a tool that highlights where this DNA “junk” is, and I demonstrated that it is widespread in engineered plasmid constructs.